How We Built Lung Cancer Screening Software for Global Clinical Use
By Domenic DiNatale
Lung cancer screening software is software a radiologist actually has to read scans on. Every shortcut you take in the architecture eventually becomes a reason a busy clinician at Mount Sinai or a screening site in Ethiopia stops trusting the tool. We've been working on this problem for a long time — the platform we now call ScreeningPlus is the third generation of a system whose lineage runs back to the original VA-PALS registry built for the U.S. Department of Veterans Affairs. This post is about what it actually took to bring that lineage forward into a modern, AI-augmented clinical platform that holds up in production.
The Problem We Inherited
VA-PALS was a serious system. It was also written in MUMPS, hosted on-premise, and shaped by the assumptions of a single VA deployment. Every assumption baked into the schema — single-tenant, one workflow, one regulatory environment — became a barrier the moment the program tried to expand. The clinical model was sound. The software substrate had become the limiting factor.
When Accumetra and Paraxial set out to build the next-generation lung cancer screening platform on top of that foundation, they came to us because we had built the original modern web layer on top of VA-PALS years earlier, integrating with its MUMPS backend through carefully bounded interfaces. We knew where the bodies were buried. More importantly, we knew which clinical behaviors of the original system had to survive any rewrite intact — and which ones were artifacts of the substrate that should be left behind.
The brief was unambiguous: rebuild the system for cloud deployment and multi-institution scale, integrate AI-assisted nodule detection directly into the radiologist's workflow, and do it without breaking the clinical fidelity that programs in screening trust to bet patient outcomes on.
Six Components, One Reading Experience
A lot of people, when they hear "lung cancer screening software," picture a single application. The reality of clinical imaging is that you are integrating with a hospital's PACS, you are doing pixel-level computer vision on volumetric CT data, you are presenting structured findings back to a radiologist in a viewer that has to handle multi-gigabyte studies smoothly, and you are persisting clinical data in a way that survives audit. No single application does all of those things well. ScreeningPlus is six components that present as one tool to the user:
- Screening+ API — a NestJS 10 / TypeScript backend with MongoDB, owning the clinical data model: CT evaluations, nodule grids, AI ingestion endpoints, patient management.
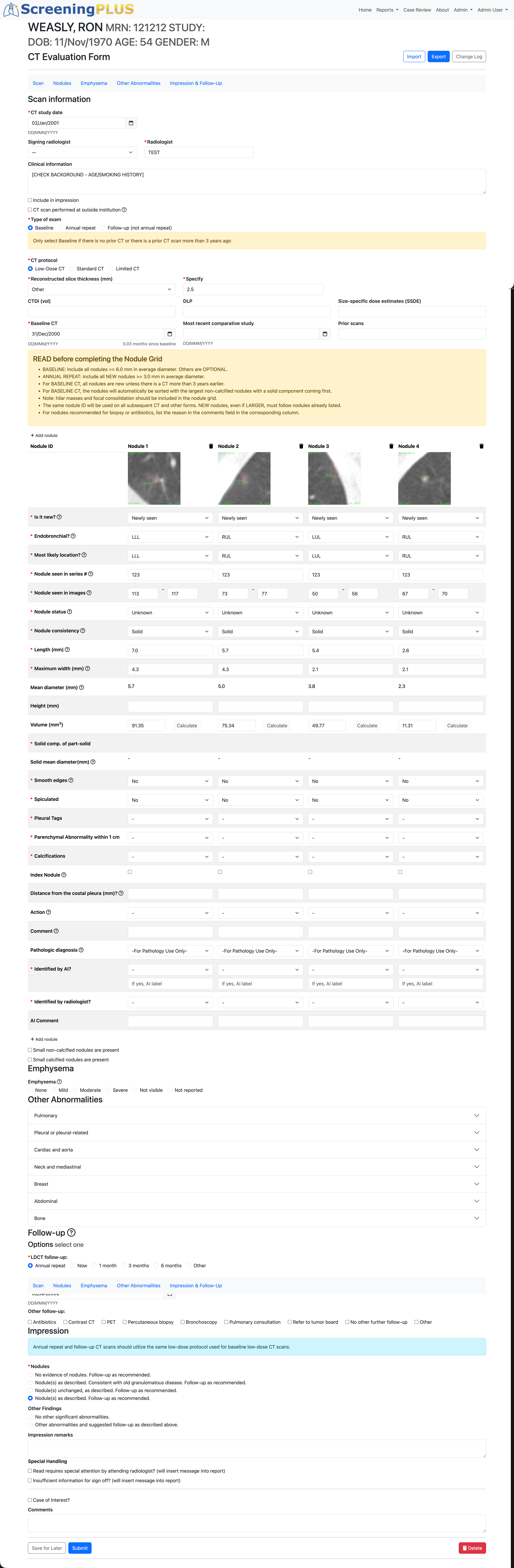
- Screening+ UI — an Angular 17 frontend where radiologists do their structured CT reads, fill out evaluation forms, work the nodule grid, and launch the viewer.
- Viewer — a React 18 application built on OHIF v3.12 with Cornerstone.js and VTK.js, doing the actual DICOM rendering, annotation, and longitudinal comparison.
- Custom AI Model — a Python image-analysis pipeline running PyTorch with nnU-Net and nnDetection for lung lobe segmentation and nodule detection, plus 3D Slicer for measurement.
- DICOM Server — a Python / Orthanc / PostgreSQL system that receives DICOM C-STORE from hospital PACS, orchestrates the AI pipeline through HTCondor, and writes results back to the rest of the platform.
Each component is independently deployable, independently scalable, and independently testable. That sounds like routine microservices advice until you remember that each one also has to integrate with hospital infrastructure that was designed around the assumption that medical imaging software is a single application sitting on a workstation. Every interface between these components is a place we had to make a deliberate decision: hospital PACS gets DICOM C-STORE; the viewer talks to Orthanc over WADO-RS and QIDO-RS; the AI pipeline gets jobs scheduled through HTCondor; everything else is HTTP.
The MUMPS-to-TypeScript Migration
Most of the rewrite was not the AI part. It was disentangling clinical logic that had been encoded in MUMPS for decades and re-expressing it in a strongly-typed, modern codebase. We did this incrementally, with one rule: clinical behavior had to be testable, not just translated. Every block of MUMPS that had been doing real work — nodule grading rules, follow-up scheduling logic, Lung-RADS categorization — got rewritten into TypeScript with a test that asserted the new code produced the same answer the old code did against the same input.
This is unglamorous work. It is also the work that decides whether a clinical migration survives contact with users. We built a corpus of historical cases out of de-identified VA-PALS data, ran both systems against that corpus, and refused to ship anything where the new system disagreed with the old until we could explain why and demonstrate the new behavior was correct. That posture — every divergence is a defect until proven otherwise — is what lets a clinical team trust a re-platforming.
How AI Actually Slots In
The most-asked question about ScreeningPlus is some variant of "is the AI making the diagnosis?" The answer is no, and the architecture makes that explicit. The AI is a batch process, scheduled by the DICOM Server through HTCondor when a CT study arrives. It runs in a Docker container on a GPU worker, takes the volumetric image data, and produces:
- Lung lobe segmentations as NIfTI volumes
- Nodule detections with epicenter coordinates and bidirectional measurements
- Pre-rendered screenshots showing detected nodules in context
- A structured JSON output the rest of the platform can consume
When the radiologist opens the case in Screening+, the AI's findings are available as a starting point. They appear in the nodule grid as importable suggestions. The radiologist can accept them, modify them, reject them, or add nodules the AI missed. Every action is logged. The clinical record reflects what the radiologist signed off on, not what the AI proposed. The viewer launches with the AI epicenters available as bookmarks; the clinician decides what is real.
This separation matters. In a regulated screening program, a tool that looks like it is making decisions on behalf of the radiologist is a different regulatory animal than a tool that surfaces findings for review. We architected toward the latter on purpose, and the data model and UI reflect that choice. The AI is an informed assistant. The radiologist is the reader.
Deploying to Three Continents
ScreeningPlus is in active clinical use at Mount Sinai Hospital and at multiple sites in Ethiopia, with continued expansion into other global lung cancer screening programs. Each of those deployments has different infrastructure realities. Mount Sinai has bandwidth, GPU resources, and well-established PACS integration. International sites often have intermittent connectivity, no on-site GPU capacity, and PACS systems with quirks the standard would not predict.
We designed for that gradient from the beginning. The DICOM Server can run on-premise; AI workloads can either run on-site GPUs or get routed through Paraxial's network infrastructure to centralized GPU resources. Hybrid configurations — on-premise data with off-premise compute — are first-class deployments, not workarounds. JWT-based authentication and encrypted MongoDB storage mean every deployment can be scoped to its own institution's data without architectural surgery.
The result is that a screening program in Addis Ababa runs the same software a screening program at Mount Sinai runs, with the same clinical workflow, against the same audit guarantees, even though the underlying infrastructure looks nothing alike. That portability is the thing that lets clinical research validation work at the scale these programs need.
What This Took
ScreeningPlus is the kind of system that doesn't reward shortcuts. The MUMPS-to-TypeScript migration was years of patient work that produced almost no externally-visible feature. The DICOM Server's interaction with HTCondor was weeks of debugging GPU job scheduling for cases nobody had run before. The viewer's longitudinal comparison flow — letting a radiologist see a nodule across two studies six months apart — required careful work in coordinate-system transforms that almost nobody outside medical imaging thinks about. None of that is the part vendors put on a slide. All of it is the part that decides whether a system gets used.
If you are building lung cancer screening software, or any clinical imaging platform with serious AI integration, the parts that look mundane in the architecture diagram — the structured-report ingestion, the audit log, the migration test corpus, the deployment portability — are the parts that determine whether the system holds up. We learned that on VA-PALS. We applied it on ScreeningPlus. We will apply it on whatever comes next.
Related work from our team:
The full project narrative lives on the ScreeningPlus case study page. For context on how we think about AI as a structural component rather than a magic black box, see What AI Actually Changes About Security — and What It Doesn't and AI Systems Have an Architecture Problem Too.